IBM announced plans to give the National Institutes of Health a database of more than 2.4 million chemical compounds. NIH will add this information to PubChem, a freely available database of chemical structures of small organic molecules and information on their biological activities.
Data mining drug discovery
The chemical compound data was pulled from about 4.7 million patents and 11 million biomedical journal abstracts from 1976-2000. According to IBM, the newly compiled data will help researches more easily visualize important relationships among chemical compounds and aid in drug discovery.
The database created by IBM was done in collaboration with AstraZeneca, Bristol-Myers Squibb, DuPont, and Pfizer. The compound data was extracted using IBM's strategic IP insight platform, a combination of data and analytics delivered by the IBM SmartCloud. The platform uses techniques such as automatic image analysis and enhanced optical recognition of chemical images and symbols.

Duke materials genome covers thousands of compounds
A materials genome repository developed by Duke University engineers will allow scientists to stop using trial-and-error methods for combining different elements to create the most efficient alloys for a promising method of producing electricity. The database is free and open to all (aflowlib.org).

These thermoelectric materials produce electricity by taking advantage of temperature differences and are currently being used in such applications as deep space satellites to campsite coolers. In the past, scientists have not had a rational basis for combining different elements to produce these energy-producing materials.
The project developed by the Duke engineers covers thousands of compounds and provides detailed "recipes" for creating the most efficient combinations for a particular purpose, much like hardware stores mix different colors to achieve the desired paint color.
"Scientists will now have a more rational basis when they decide which elements to combine for their thermoelectric devices," said Stefano Curtarolo, associate professor of mechanical engineering and materials sciences at Duke's Pratt School of Engineering.

BIGPLANT's evolutionary relationships of 150 plant species
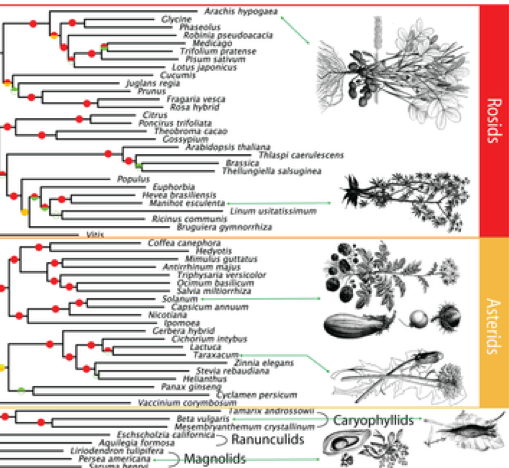
Scientists at New York University's Center for Genomics and Systems Biology have created the largest genome-based tree of life for seed plants to date. Their findings, published in the journal PLoS Genetics, plot the evolutionary relationships of 150 different species of plants based on advanced genome-wide analysis of gene structure and function. The complete phylogenomic matrix and trees are available at the BIGPLANT (http://nypg.bio.nyu.edu) website.
This allows scientists to reconstruct the pattern of events that led to the vast number of plant species and could help identify genes used to improve seed quality for agriculture. The sequences of the plants' genomes--all of the biological information needed to build and maintain an organism, encoded in DNA--were either culled from pre-existing databases or generated in the field and at the New York BotanicalGarden from live specimens.
"Previously, phylogenetic trees were constructed from standard sets of genes and were
used to identify the relationships of species," explains Gloria Coruzzi, professor of biology at New York University. "In our novel approach, we create the phylogeny based on all the genes in a genome, and then use the phylogeny to identify which genes provide positive support for the divergence of species."
The data and software resources generated by the researchers are publicly available. These studies could have implications for improving the quality of seeds and, in turn, agricultural products ranging from food to clothing.


